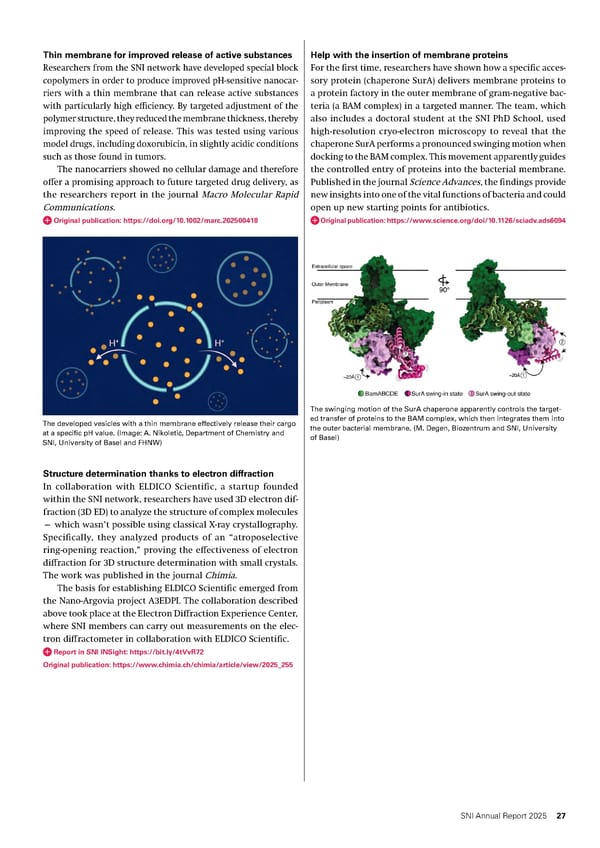
Help with the insertion of membrane proteins For the first time, researchers have shown how a specific acces- sory protein (chaperone SurA) delivers membrane proteins to a protein factory in the outer membrane of gram-negative bac- teria (a BAM complex) in a targeted manner. The team, which also includes a doctoral student at the SNI PhD School, used high-resolution cryo-electron microscopy to reveal that the chaperone SurA performs a pronounced swinging motion when docking to the BAM complex. This movement apparently guides the controlled entry of proteins into the bacterial membrane. Published in the journal Science Advances, the findings provide new insights into one of the vital functions of bacteria and could open up new starting points for antibiotics. Original publication: https://www.science.org/doi/10.1126/sciadv.ads6094 The swinging motion of the SurA chaperone apparently controls the target- ed transfer of proteins to the BAM complex, which then integrates them into the outer bacterial membrane. (M. Degen, Biozentrum and SNI, University of Basel) Thin membrane for improved release of active substances Researchers from the SNI network have developed special block copolymers in order to produce improved pH-sensitive nanocar- riers with a thin membrane that can release active substances with particularly high efficiency. By targeted adjustment of the polymer structure, they reduced the membrane thickness, thereby improving the speed of release. This was tested using various model drugs, including doxorubicin, in slightly acidic conditions such as those found in tumors. The nanocarriers showed no cellular damage and therefore offer a promising approach to future targeted drug delivery, as the researchers report in the journal Macro Molecular Rapid Communications. Original publication: https://doi.org/10.1002/marc.202500418 The developed vesicles with a thin membrane effectively release their cargo at a specific pH value. (Image: A. Nikoleti ć , Department of Chemistry and SNI, University of Basel and FHNW) Structure determination thanks to electron diffraction In collaboration with ELDICO Scientific, a startup founded within the SNI network, researchers have used 3D electron dif- fraction (3D ED) to analyze the structure of complex molecules — which wasn’t possible using classical X-ray crystallography. Specifically, they analyzed products of an “atroposelective ring-opening reaction,” proving the effectiveness of electron diffraction for 3D structure determination with small crystals. The work was published in the journal Chimia. The basis for establishing ELDICO Scientific emerged from the Nano-Argovia project A3EDPI. The collaboration described above took place at the Electron Diffraction Experience Center, where SNI members can carry out measurements on the elec- tron diffractometer in collaboration with ELDICO Scientific. Report in SNI INSight: https://bit.ly/4tVvR72 Original publication: https://www.chimia.ch/chimia/article/view/2025_255 27 SNI Annual Report 2025
 Annual Report 2025 - Swiss Nanoscience Institute Page 26 Page 28
Annual Report 2025 - Swiss Nanoscience Institute Page 26 Page 28